Quick Start¶
Get started with qpx in minutes - from installation to your first data conversion.
Quick Start Flow¶
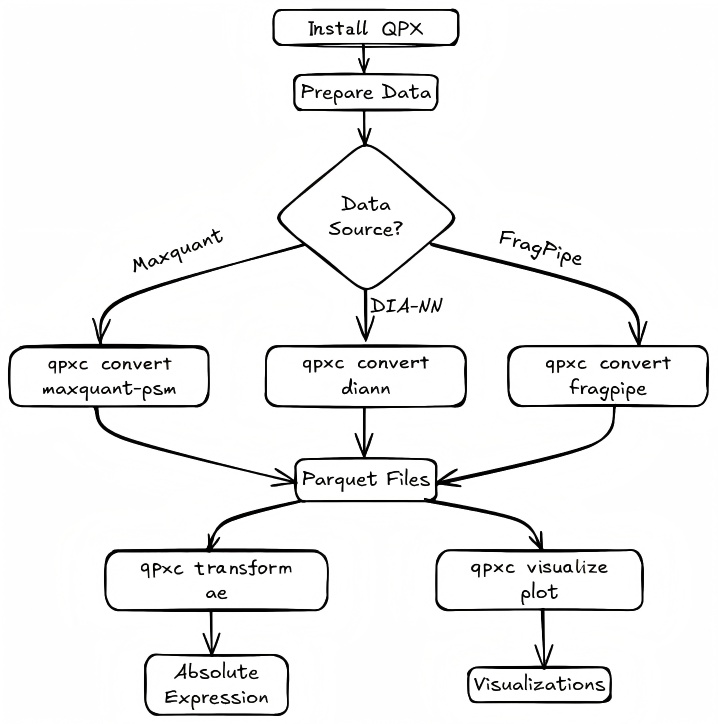
Prerequisites¶
Before installing qpx, ensure you have:
- Python 3.10 or higher - Check with
python --version - pip - Python package manager (included with Python)
- Optional: conda/mamba for environment management
Installation¶
Verify Installation¶
After installation, verify qpx is working correctly:
You should see output similar to:
Usage: qpxc [OPTIONS] COMMAND [ARGS]...
qpxc -- quantitative proteomics data format tools.
Options:
--version Show the version and exit.
-h, --help Show this message and exit.
Commands:
convert Convert external tool outputs to QPX format.
info Show information about a QPX dataset.
ontology Manage CV ontology data (PSI-MS, PRIDE CV).
query Query and inspect QPX datasets.
transform Transform QPX data into derived representations.
validate Validate a QPX dataset or structure against the canonical...
Your First Conversion¶
Let's convert some sample MaxQuant data to QPX format.
Step 1: Download Sample Data¶
# Create a working directory
mkdir qpx-tutorial && cd qpx-tutorial
# Download sample MaxQuant msms.txt file
curl -L -o msms.txt \
"https://raw.githubusercontent.com/bigbio/qpx/main/tests/examples/maxquant/maxquant_simple/msms.txt"
Step 2: Convert to QPX Format¶
# Convert MaxQuant PSM data to QPX parquet format
qpxc convert maxquant \
--msms-file msms.txt \
--output-folder ./output \
--structures psm \
--verbose
Step 3: Verify the Output¶
Step 4: Inspect the Data (Optional)¶
# Using Python to read the parquet file
import pyarrow.parquet as pq
table = pq.read_table("output/psm-*.psm.parquet")
df = table.to_pandas()
print(f"Total PSMs: {len(df)}")
print(f"Columns: {list(df.columns)}")
print(df.head())
What's Next?¶
Now that you've completed your first conversion, explore more:
| Next Step | Description |
|---|---|
| Examples Overview | More conversion and analysis examples |
| Convert Commands | All available data converters |
| Transform Commands | Data transformation tools |
| Format Specification | Understanding QPX data formats |
Common Commands¶
# Convert DIA-NN data
qpxc convert diann --report-path report.tsv --sdrf-file data.sdrf.tsv --output-folder ./output
# Convert FragPipe data
qpxc convert fragpipe --psm-file psm.tsv --output-folder ./output
# Validate a QPX dataset
qpxc validate --dataset-folder ./output
# Query a QPX dataset
qpxc query --dataset-folder ./output --sql "SELECT * FROM psm LIMIT 10"
Need Help?¶
- Run
qpxc <command> --helpfor detailed command help - Check the Troubleshooting guide
- Visit our GitHub Issues for support